Replicate Weights in the American Community Survey / Puerto Rican Community Survey
Go Back to IPUMS-USA Documentation
Summary
- Why might I want to use replicate weights?
- What are replicate weights?
- Does using replicate weights make any substantive difference?
- How do I obtain replicate standard errors from IPUMS-USA data?
- Is there any way to do this automatically in major statistical packages?
- Can I simply divide the full sample into 80 random subsamples from the full sample and calculate replicate standard errors manually?
- How are the ACS/PRCS replicate weights calculated?
Why might I want to use replicate weights?
Replicate weights allow researchers to obtain confidence intervals and run significance tests for estimates that parallel the balanced half-sample procedure used by statisticians at the Census Bureau. By dividing surveyed strata into multiple balanced half-samples, a single sample is made to simulate a range of possible alternative samples, generating more informed standard error estimates that mimic the theoretical basis of standard errors while retaining all information about the complex sample design.
What are replicate weights?
Calculating standard errors with balanced half-samples requires knowledge of survey design parameters, specifically the Primary Sampling Unit (PSU) and stratum for each respondent. However, these are typically kept confidential to safeguard individuals' identities. Replicate weights solve this dilemma by containing the necessary information to derive the Census Bureau’s standard error calculations without requiring access to PSU or strata identifiers. Each set of weights represents a balanced half-sample created by the Census Bureau using Successive Difference Replication, allowing researchers to replicate calculations without compromising the identity of respondents.
Replicate weights are currently available for the 2005-onward American Community Survey and Puerto Rican Community Survey data. In the ACS and PRCS, there are 80 separate replicate weights at the household and person levels.
Does using replicate weights make any substantive difference?
In IPUMS testing of ACS/PRCS data, replicate weights usually increase standard errors. This increase is generally not large enough to alter the significance level of coefficients, though marginally significant coefficients may become clearly nonsignificant. The more obvious effect of using replicate weights is on the width of confidence intervals, which can change substantially.
How do I obtain replicate standard errors from IPUMS-USA data?
There are 3 main steps:
- Run your analysis using the full-sample weights (PERWT and HHWT are the main IPUMS-USA weights). Record the statistic you are interested in (e.g., the mean income of veterans, or the coefficient describing the relationship between income and whether one has health insurance coverage).
- Run your analysis again using each set of replicate weights. First, run the analysis using REPWTP1, then again using REPWTP2, then again using REPWTP3, and so on up to the final set of replicate weights. After each set, record the statistic you are interested in. (N.B.: If you are analyzing a household-only file, be sure to use REPWT1, REPWT2, etc.)
-
Insert the above results into the following formula:
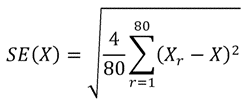
where X is the result from the analysis using the full-sample weight and Xr is the result from the analysis using the r-th set of replicate weights.
Is there any way to do this automatically in major statistical packages?
Yes. Although the replicate weights contained in the IPUMS-USA data are calculated using the successive difference replication method, there are multiple equivalent methods for implementing these as survey design parameters.
R
To use IPUMS-USA replicate weights in R, you must use the srvyr package.
install.packages("srvyr")
library("srvyr")Next, you'll create a survey object using the replicate weights.
svy <- as_survey_rep(data, weight = PERWT,
repweights = matches("REPWTP[0-9]+"),
type = "ACS", mse = TRUE)Any calculations you'd like to make with the replicate weights should be done with the object 'svy' instead of the object 'data'.
svy %>%
group_by(RACE)%>%
summarize(mean_age = survey_mean((AGE), vartype="ci"))Stata
To use IPUMS-USA replicate weights in Stata, you must first svyset the data.
. svyset[pweight=perwt], vce(sdr) sdrweight(repwtp1-repwtp80) dof(79) mseEarlier versions of Stata (versions 11.0 and before) can also handle successive difference replicate weights. Correspondence with StataCorp statisticians and IPUMS testing revealed that successive difference replicate weights can be treated as Jackknife replicate weights if the options are specified correctly.
. svyset [pw=perwt], jkrweight(repwtp1-repwtp80, multiplier(.05)) ///
vce(jackknife) dof(79) mse- The sample should be treated as a single stratum (the weights contain the relevant information from the sample design), so no PSU should be specified.
- The full-sample weight must be specified.
- Specifying the replicate weight variable list with a wildcard character (
repwtp*) rather than with a range of variables (repwtp1-repwtp80) will not produce correct results because IPUMS-USA data contain a variable called REPWTP, which merely indicates the presence of replicate weights and is coded 1 for every case. - The
fpc()suboption should not be specified. - The
dof()suboption should be specified to calculate confidence errors for estimates with 79 degrees of freedom (80 replicates - 1). - There are a few differences between the SDR and Jackknife specifications.
- With SDR, You must specify the
vce(sdr)and themseoptions. - With Jackknife replication,
multiplier()suboption must be specified with the quotient from the formula (4/80 = 0.05). If you are not using ACS/PRCS data and have a different number of replicate weights, you will need to adjust the multiplier accordingly.
After
svysettingthe data, you run the command using thesvy:prefix, which passes along the options you defined above.. svy: commandStata will execute this command using the full-sample weights and again for each set of replicate weights. There are two important things to note:
- Not all Stata commands can be run with the
svy:prefix. Type. help svy_estimationto see a list of valid commands. -
If you want to limit your replicate analyses to a subset of the sample (for example, all persons aged 25-64 or all African Americans), you should not use if or in. Instead, use the
subpop()option before the colon, as in
Note that you must first define the subpopulation with a dichotomous variable coded 0 for all cases that should be excluded from the analysis. See this page for a helpful discussion of. gen byte age25_64 = age>=25 & age<=64 . svy, subpop(age25_64): commandsubpop()nuances.
SAS
In SAS, instead of declaring a set of survey options all at once, you must declare them in each statistical procedure. Replicate weights can be applied in any statistical procedure that accepts the jackknife method of variance estimation by using the JKCOEFS option to set the coefficients for the jackknife replicate weights to be 0.05 (the result of 4/80 in the formula above). In regression, for example, type:
PROC SURVEYREG data=dataset VARMETHOD=jackknife; MODEL dependent_variable=independent_variable(s) WEIGHT perwt; REPWEIGHTS repwtp1-repwtp80 /JKCOEFS=0.05; run;See also the Census Bureau's "Estimating ASEC Variances with Replicate Weights" document for sample SAS code that can be adapted to calculate replicate standard errors manually. (Although this document describes replicate weights in the Current Population Survey, the material on using replicate weights applies to ACS/PRCS replicate weights as well, with some adaptation.)
SPSS
As of September 2011, SPSS (version 19.0) cannot handle successive difference replicate weights. SPSS does not allow for replicate-based variance estimation unless it performs the resampling itself.
Can I simply divide the full sample into 80 random subsamples from the full sample and calculate replicate standard errors manually?
No. Replicate weights contain full information about the complex sample design of the ACS/PRCS, and this information would be lost when drawing random subsamples. Furthermore, replicate samples incorporate information from all cases in the full sample. In contrast, random subsamples would each be 1/80th the size of a single replicate subsample.
How are the ACS/PRCS replicate weights calculated?
As mentioned, replicate weights in the ACS and PRCS are constructed using the successive difference replication method. This involves creating a k x k Hadamard matrix (where k is the number of replicate weights desired), assigning sample cases to rows in the matrix and calculating a replicate factor from the row values, and finally multiplying the full-sample weight by these replicate factors. For more details, see the Census Bureau's "Estimating ASEC Variances with Replicate Weights" document, written for the CPS, as well as the following:
- Fay, Robert, and George Train. 1995. "Aspects of Survey and Model-Based Postcensal Estimation of Income and Poverty Characteristics for States and Counties." Proceedings of the Section on Government Statistics, American Statistical Association, Alexandria, VA, pp. 154-159. (pdf)
- Wolter, Kirk. 2007. Introduction to Variance Estimation, 2nd ed. New York: Springer.
- With SDR, You must specify the